
We retrieved 159 OA-related per-view publications from the Web of Science (a list of publicaitons used by this release see below) and read and collected genes associated OA reported by the publications. At least one evidence was found to support the corresponding gene as a biomarker for OA: genome-wide association study (GWAS), differential expressed gene (DEG) or differential methylation region (DMR) analysis (Boer et al., 2021; Ramos et al. 2014; Dunn et al., 2019; Ali et al., 2020; Ji et al., 2018).
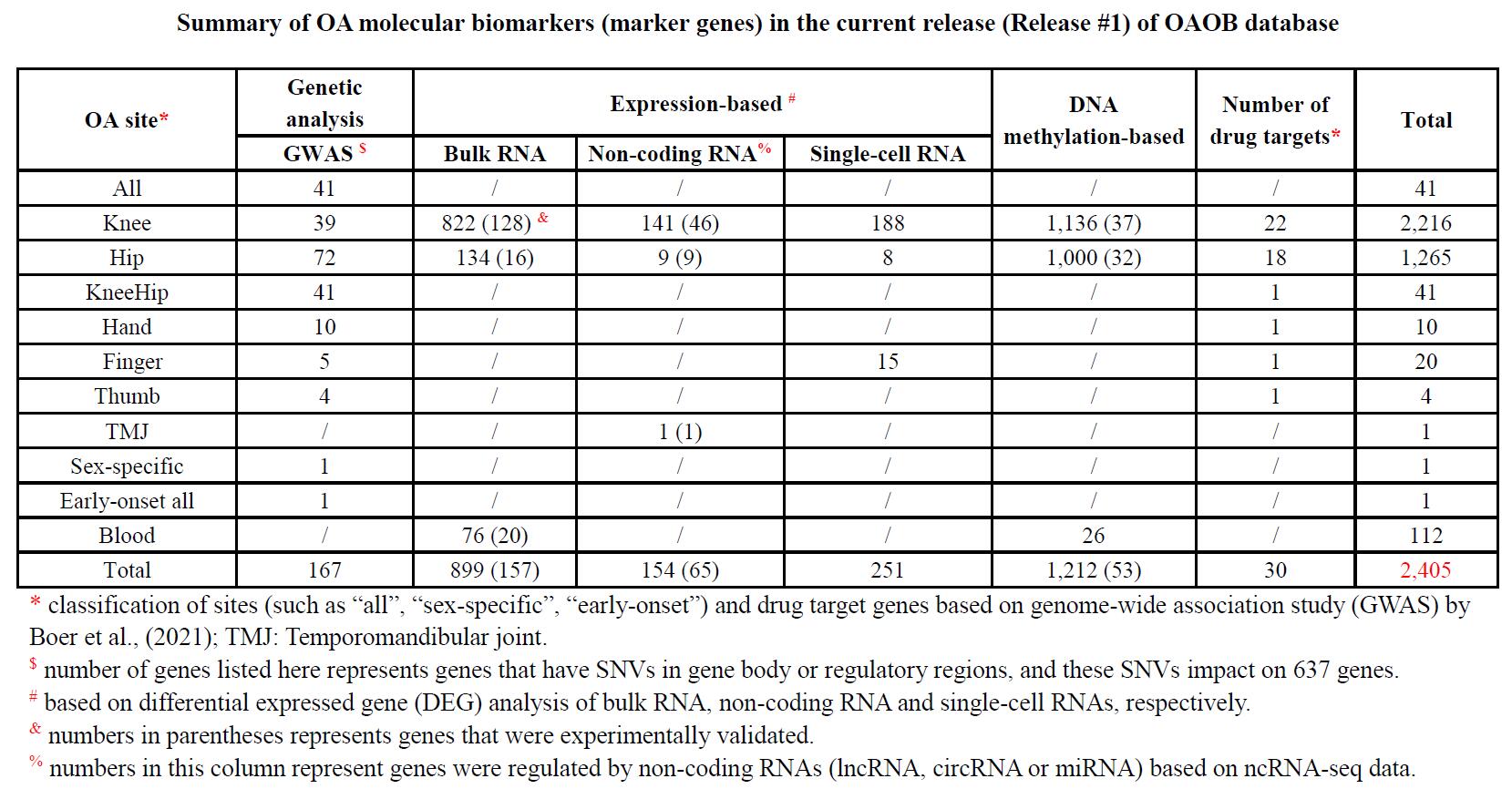
As a results, a total of 2,405 OA marker genes which have been identified from different joints were collected (Table 1). The 2,405 OA marker genes have diverse evidences, such as 167 by GWAS, 899 by DEG analysis based on transcriptomic (bull RNA, 65 were further experimentally validated) and 1,212 by DMR analysis (56 further validated). In the 167 marker genes or OA-associated effector genes by GWAS, most 126 (17.5%) were also included in the gene set independently identified by DEG or DMR analysis.
DEG analysis base on small RNAs and single-cell RNAs (scRNA), also found several hundreds of mark genes, including 154 genes coding genes that were regulated by 176 non-coding genes (11 lncRNAs, five circRNAs and 160 miRNAs), and 251 genes identified by scRNA-seq data.
With more clinical significance, 112 OA marker genes (22 experimentally validated) were identified from blood of OA patients, which had significantly different expression or methylation level relative to normal (Table 1).
Additionally, a total of 30 OA marker genes can be targeted by available drugs based on DrugBank database, which provide another useful clinical approach to cure OA.
The above marker genes are predictive of OA progression and valuable diagnosis biomarkers with superior ability to measure early and subtle changes in OA progression compared to traditional radiographic measures.
(Qinjie Chu, Lingjuan Xie, Xiaoxu Xu, Xiaotian Yang, Longjiang Fan)
1. Ahn J, Kumar H, Cha BH, et al. AIMP1 downregulation restores chondrogenic characteristics of dedifferentiated/degenerated chondrocytes by enhancing TGF-β signal. Cell Death Dis. 2016;7(2):e2099. doi:10.1038/cddis.2016.17
2. Akhtar N, Rasheed Z, Ramamurthy S, Anbazhagan AN, Voss FR, Haqqi TM. MicroRNA-27b regulates the expression of matrix metalloproteinase 13 in human osteoarthritis chondrocytes. Arthritis Rheum. 2010;62(5):1361-1371. doi:10.1002/art.27329
3. Ali SA, Gandhi R, Potla P, et al. Sequencing identifies a distinct signature of circulating microRNAs in early radiographic knee osteoarthritis. Osteoarthritis Cartilage. 2020;28(11):1471-1481. doi:10.1016/j.joca.2020.07.003
4. Alnajjar FA, Sharma-Oates A, Wijesinghe SN, et al. The Expression and Function of Metastases Associated Lung Adenocarcinoma Transcript-1 Long Non-Coding RNA in Subchondral Bone and Osteoblasts from Patients with Osteoarthritis. Cells. 2021;10(4):786. doi:10.3390/cells10040786
5. Alvarez-Garcia O, Fisch KM, Wineinger NE, et al. Increased DNA Methylation and Reduced Expression of Transcription Factors in Human Osteoarthritis Cartilage. Arthritis Rheumatol. 2016;68(8):1876-1886. doi:10.1002/art.39643
6. An Y, Wan G, Tao J, Cui M, Zhou Q, Hou W. Down-regulation of microRNA-203a suppresses IL-1β-induced inflammation and cartilage degradation in human chondrocytes through Smad3 signaling. Biosci Rep. 2020;40(3):BSR20192723. doi:10.1042/BSR20192723
7. arcOGEN Consortium, arcOGEN Collaborators. Identification of new susceptibility loci for osteoarthritis (arcOGEN): a genome-wide association study. The Lancet. 2012; 380(9844): 815-823. doi:10.1016/S0140-6736(12)60681-3
8. Aref-Eshghi E, Liu M, Razavi-Lopez SB, et al. SMAD3 Is Upregulated in Human Osteoarthritic Cartilage Independent of the Promoter DNA Methylation. J Rheumatol. 2016;43(2):388-394. doi:10.3899/jrheum.150609
9. Aref-Eshghi E, Zhang Y, Liu M, et al. Genome-wide DNA methylation study of hip and knee cartilage reveals embryonic organ and skeletal system morphogenesis as major pathways involved in osteoarthritis. BMC Musculoskelet Disord. 2015;16:287. doi:10.1186/s12891-015-0745-5
10. Bai Y, Chen K, Zhan J, Wu M. miR-122/SIRT1 axis regulates chondrocyte extracellular matrix degradation in osteoarthritis. Biosci Rep. 2020;40(6):BSR20191908. doi:10.1042/BSR20191908
11. Bai Z M, Kang M M, Zhou X F, et al. CircTMBIM6 promotes osteoarthritis-induced chondrocyte extracellular matrix degradation via miR-27a/MMP13 axis. Eur Rev Med Pharmacol Sci. 2020;24(15): 7927-7936. doi:10.26355/eurrev_202008_22475
12. Boer CG, Hatzikotoulas K, Southam L, et al. Deciphering osteoarthritis genetics across 826,690 individuals from 9 populations. Cell. 2021;184(18):4784-4818.e17. doi:10.1016/j.cell.2021.07.038
13. Bomer N, den Hollander W, Ramos YF, et al. Underlying molecular mechanisms of DIO2 susceptibility in symptomatic osteoarthritis. Ann Rheum Dis. 2015;74(8):1571-1579. doi:10.1136/annrheumdis-2013-204739
14. Bonin CA, Lewallen EA, Baheti S, et al. Identification of differentially methylated regions in new genes associated with knee osteoarthritis. Gene. 2016;576(1 Pt 2):312-318. doi:10.1016/j.gene.2015.10.037
15. Cao J, Liu Z, Zhang L, Li J. miR-940 regulates the inflammatory response of chondrocytes by targeting MyD88 in osteoarthritis. Mol Cell Biochem. 2019;461(1-2):183-193. doi:10.1007/s11010-019-03601-z
16. Cao L, Wang Y, Wang Q, Huang J. LncRNA FOXD2-AS1 regulates chondrocyte proliferation in osteoarthritis by acting as a sponge of miR-206 to modulate CCND1 expression. Biomed Pharmacother. 2018;106:1220-1226. doi:10.1016/j.biopha.2018.07.048
17. Cao X, Duan Z, Yan Z, et al. miR-195 contributes to human osteoarthritis via targeting PTHrP. J Bone Miner Metab. 2019;37(4):711-721. doi:10.1007/s00774-018-0973-5
18. Casalone E, Tachmazidou I, Zengini E, et al. A novel variant in GLIS3 is associated with osteoarthritis. Ann Rheum Dis. 2018;77(4):620-623.
doi:10.1136/annrheumdis-2017-211848
19. Chang ZK, Meng FG, Zhang ZQ, et al. MicroRNA-193b-3p regulates matrix metalloproteinase 19 expression in interleukin-1β-induced human chondrocytes. J Cell Biochem. 2018;119(6):4775-4782. doi:10.1002/jcb.26669
20. Chen J, Wu X. MicroRNA-103 contributes to osteoarthritis development by targeting Sox6. Biomed Pharmacother. 2019;118:109186. doi:10.1016/j.biopha.2019.109186
21. Chen L, Li Q, Wang J, et al. MiR-29b-3p promotes chondrocyte apoptosis and facilitates the occurrence and development of osteoarthritis by targeting PGRN. J Cell Mol Med. 2017;21(12):3347-3359. doi:10.1111/jcmm.13237
22. Chen YJ, Chang WA, Wu LY, Huang CF, Chen CH, Kuo PL. Identification of Novel Genes in Osteoarthritic Fibroblast-Like Synoviocytes Using Next-Generation Sequencing and Bioinformatics Approaches. Int J Med Sci. 2019;16(8):1057-1071. doi:10.7150/ijms.35611
23. Cheng F, Hu H, Sun K, Yan F, Geng Y. miR-455-3p enhances chondrocytes apoptosis and inflammation by targeting COL2A1 in the in vitro osteoarthritis model. Biosci Biotechnol Biochem. 2020;84(4):695-702. doi:10.1080/09168451.2019.1690974
24. Cheng W, Hao C Y, Zhao S, et al. SNHG16 promotes the progression of osteoarthritis through activating microRNA-93-5p/CCND1 axis. Eur Rev Med Pharmacol Sci. 2019;23(21): 9222-9229. doi:10.26355/eurrev_201911_19414
25. Cheung KS, Hashimoto K, Yamada N, Roach HI. Expression of ADAMTS-4 by chondrocytes in the surface zone of human osteoarthritic cartilage is regulated by epigenetic DNA de-methylation. Rheumatol Int. 2009;29(5):525-534. doi:10.1007/s00296-008-0744-z
26. Chou CH, Jain V, Gibson J, et al. Synovial cell cross-talk with cartilage plays a major role in the pathogenesis of osteoarthritis. Sci Rep. 2020;10(1):10868. doi:10.1038/s41598-020-67730-y
27. Chou CH, Wu CC, Song IW, et al. Genome-wide expression profiles of subchondral bone in osteoarthritis. Arthritis Res Ther. 2013;15(6):R190. doi:10.1186/ar4380
28. Dai Y, Liu S, Xie X, Ding M, Zhou Q, Zhou X. MicroRNA 31 promotes chondrocyte proliferation by targeting C X C motif chemokine ligand 12. Mol Med Rep. 2019;19(3):2231-2237. doi:10.3892/mmr.2019.9859
29. Dang X, Lian L, Wu D. The diagnostic value and pathogenetic role of lncRNA-ATB in patients with osteoarthritis. Cell Mol Biol Lett. 2018;23:55. doi:10.1186/s11658-018-0118-9
30. Day-Williams AG, Southam L, Panoutsopoulou K, et al. A variant in MCF2L is associated with osteoarthritis. Am J Hum Genet. 2011;89(3):446-450. doi:10.1016/j.ajhg.2011.08.001
31. Dehne T, Karlsson C, Ringe J, Sittinger M, Lindahl A. Chondrogenic differentiation potential of osteoarthritic chondrocytes and their possible use in matrix-associated autologous chondrocyte transplantation. Arthritis Res Ther. 2009;11(5):R133. doi:10.1186/ar2800
32. Del Rey MJ, Usategui A, Izquierdo E, et al. Transcriptome analysis reveals specific changes in osteoarthritis synovial fibroblasts. Ann Rheum Dis. 2012;71(2):275-280. doi:10.1136/annrheumdis-2011-200281
33. den Hollander W, Boer CG, Hart DJ, et al. Genome-wide association and functional studies identify a role for matrix Gla protein in osteoarthritis of the hand. Ann Rheum Dis. 2017;76(12):2046-2053. doi:10.1136/annrheumdis-2017-211214
34. den Hollander W, Ramos YF, Bos SD, et al. Knee and hip articular cartilage have distinct epigenomic landscapes: implications for future cartilage regeneration approaches. Ann Rheum Dis. 2014;73(12):2208-2212. doi:10.1136/annrheumdis-2014-205980
35. Dicks A, Wu CL, Steward N, Adkar SS, Gersbach CA, Guilak F. Prospective isolation of chondroprogenitors from human iPSCs based on cell surface markers identified using a CRISPR-Cas9-generated reporter. Stem Cell Res Ther. 2020;11(1):66. doi:10.1186/s13287-020-01597-8
36. Dou P, Hu R, Zhu W, et al. Long non-coding RNA HOTAIR promotes expression of ADAMTS-5 in human osteoarthritic articular chondrocytes. Pharmazie. 2017;72(2):113-117. doi:10.1691/ph.2017.6649
37. Duan L, Duan D, Wei W, et al. MiR-19b-3p attenuates IL-1β induced extracellular matrix degradation and inflammatory injury in chondrocytes by targeting GRK6. Mol Cell Biochem. 2019;459(1-2):205-214. doi:10.1007/s11010-019-03563-2
38. Duan ZX, Huang P, Tu C, et al. MicroRNA-15a-5p Regulates the Development of Osteoarthritis by Targeting PTHrP in Chondrocytes. Biomed Res Int. 2019;2019:3904923. doi:10.1155/2019/3904923
39. El Mansouri FE, Nebbaki SS, Kapoor M, et al. Lysine-specific demethylase 1-mediated demethylation of histone H3 lysine 9 contributes to interleukin 1β-induced microsomal prostaglandin E synthase 1 expression in human osteoarthritic chondrocytes. Arthritis Res Ther. 2014;16(3):R113. doi:10.1186/ar4564
40. Endisha H, Datta P, Sharma A, et al. MicroRNA-34a-5p Promotes Joint Destruction During Osteoarthritis. Arthritis Rheumatol. 2021;73(3):426-439. doi:10.1002/art.41552
41. Evangelou E, Kerkhof HJ, Styrkarsdottir U, et al. A meta-analysis of genome-wide association studies identifies novel variants associated with osteoarthritis of the hip. Ann Rheum Dis. 2014;73(12):2130-2136. doi:10.1136/annrheumdis-2012-203114
42. Evangelou E, Valdes AM, Castano-Betancourt MC, et al. The DOT1L rs12982744 polymorphism is associated with osteoarthritis of the hip with genome-wide statistical significance in males. Ann Rheum Dis. 2013;72(7):1264-1265. doi:10.1136/annrheumdis-2012-203182
43. Evangelou E, Valdes AM, Kerkhof HJ, et al. Meta-analysis of genome-wide association studies confirms a susceptibility locus for knee osteoarthritis on chromosome 7q22. Ann Rheum Dis. 2011;70(2):349-355. doi:10.1136/ard.2010.132787
44. Fan X, Yuan J, Xie J, et al. Long non-protein coding RNA DANCR functions as a competing endogenous RNA to regulate osteoarthritis progression via miR-577/SphK2 axis. Biochem Biophys Res Commun. 2018;500(3):658-664. doi:10.1016/j.bbrc.2018.04.130
45. Fan Y, Gao D, Zhang Y, et al. Genome-Wide Differentially Methylated Region Analysis to Reveal Epigenetic Differences of Articular Cartilage in Kashin-Beck Disease and Osteoarthritis. Front Cell Dev Biol. 2021;9:636291. doi:10.3389/fcell.2021.636291
46. Fernández-Tajes J, Soto-Hermida A, Vázquez-Mosquera ME, et al. Genome-wide DNA methylation analysis of articular chondrocytes reveals a cluster of osteoarthritic patients. Ann Rheum Dis. 2014;73(4):668-677. doi:10.1136/annrheumdis-2012-202783
47. Fisch KM, Gamini R, Alvarez-Garcia O, et al. Identification of transcription factors responsible for dysregulated networks in human osteoarthritis cartilage by global gene expression analysis. Osteoarthritis Cartilage. 2018;26(11):1531-1538. doi:10.1016/j.joca.2018.07.012
48. Gao C, Pu H, Zhou Q, et al. Two reactive behaviors of chondrocytes in an IL-1β-induced inflammatory environment revealed by the single-cell RNA sequencing. Aging (Albany NY). 2021;13(8):11646-11664. doi:10.18632/aging.202857
49. Gao S, Liu L, Zhu S, et al. MicroRNA-197 regulates chondrocyte proliferation, migration, and inflammation in pathogenesis of osteoarthritis by targeting EIF4G2. Biosci Rep. 2020;40(9):BSR20192095. doi:10.1042/BSR20192095
50. Gong Y, Yang J, Li X, et al. A systematic dissection of human primary osteoblastsin vivoat single-cell resolution. 2020. doi:10.1101/2020.05.12.091975
51. Hackinger S, Trajanoska K, Styrkarsdottir U, et al. Evaluation of shared genetic aetiology between osteoarthritis and bone mineral density identifies SMAD3 as a novel osteoarthritis risk locus. Hum Mol Genet. 2017;26(19):3850-3858. doi:10.1093/hmg/ddx285
52. He D, Liu J, Hai Y, et al. Increased DOT1L in synovial biopsies of patients with OA and RA. Clin Rheumatol. 2018;37(5):1327-1332. doi:10.1007/s10067-017-3941-x
53. He J, Su X, Xie W. MiR-582-3p alleviates osteoarthritis progression by targeting YAP1. Mol Immunol. 2020;128:258-267. doi:10.1016/j.molimm.2020.10.022
54. He X, Gao K, Lu S, Wu R. LncRNA HOTTIP leads to osteoarthritis progression via regulating miR-663a/ Fyn-related kinase axis. BMC Musculoskelet Disord. 2021;22(1):67. doi:10.1186/s12891-020-03861-7
55. Hu J, Wang Z, Pan Y, et al. MiR-26a and miR-26b mediate osteoarthritis progression by targeting FUT4 via NF-κB signaling pathway. Int J Biochem Cell Biol. 2018;94:79-88. doi:10.1016/j.biocel.2017.12.003
56. Hu S, Zhao X, Mao G, et al. MicroRNA-455-3p promotes TGF-β signaling and inhibits osteoarthritis development by directly targeting PAK2. Exp Mol Med. 2019;51(10):1-13. doi:10.1038/s12276-019-0322-3
57. Iliopoulos D, Malizos KN, Tsezou A. Epigenetic regulation of leptin affects MMP-13 expression in osteoarthritic chondrocytes: possible molecular target for osteoarthritis therapeutic intervention. Ann Rheum Dis. 2007;66(12):1616-1621. doi:10.1136/ard.2007.069377
58. Jeffries MA, Donica M, Baker LW, et al. Genome-Wide DNA Methylation Study Identifies Significant Epigenomic Changes in Osteoarthritic Subchondral Bone and Similarity to Overlying Cartilage. Arthritis Rheumatol. 2016;68(6):1403-1414. doi:10.1002/art.39555
59. Ji Q, Xu X, Kang L, et al. Hematopoietic PBX-interacting protein mediates cartilage degeneration during the pathogenesis of osteoarthritis. Nat Commun. 2019;10(1):313. doi:10.1038/s41467-018-08277-5
60. Ji Q, Zheng Y, Zhang G, et al. Single-cell RNA-seq analysis reveals the progression of human osteoarthritis. Ann Rheum Dis. 2019;78(1):100-110. doi:10.1136/annrheumdis-2017-212863
61. Jia D, Li Y, Han R, et al. miR 146a 5p expression is upregulated by the CXCR4 antagonist TN14003 and attenuates SDF 1 induced cartilage degradation. Mol Med Rep. 2019;19(5):4388-4400. doi:10.3892/mmr.2019.10076
62. Jin Z, Ren J, Qi S. Human bone mesenchymal stem cells-derived exosomes overexpressing microRNA-26a-5p alleviate osteoarthritis via down-regulation of PTGS2. Int Immunopharmacol. 2020;78:105946. doi:10.1016/j.intimp.2019.105946
63. Kanamoto T, Hikida M, Sato S, et al. Integrin α2β1 plays an important role in the interaction between human articular cartilage-derived chondrocytes and atelocollagen gel. Sci Rep. 2021;11(1):1757. doi:10.1038/s41598-021-81378-2
64. Kerkhof HJ, Lories RJ, Meulenbelt I, et al. A genome-wide association study identifies an osteoarthritis susceptibility locus on chromosome 7q22. Arthritis Rheum. 2010;62(2):499-510. doi:10.1002/art.27184
65. Klinger P, Beyer C, Ekici AB, et al. The Transient Chondrocyte Phenotype in Human Osteophytic Cartilage: A Role of Pigment Epithelium-Derived Factor?. Cartilage. 2013;4(3):249-255. doi:10.1177/1947603513480809
66. Ko JY, Lee MS, Lian WS, et al. MicroRNA-29a Counteracts Synovitis in Knee Osteoarthritis Pathogenesis by Targeting VEGF. Sci Rep. 2017;7(1):3584. doi:10.1038/s41598-017-03616-w
67. Kopańska M, Szala D, Czech J, et al. MiRNA expression in the cartilage of patients with osteoarthritis [published correction appears in J Orthop Surg Res. 2017 Jun 13;12 (1):90]. J Orthop Surg Res. 2017;12(1):51. doi:10.1186/s13018-017-0542-y
68. Lai Z, Cao Y. Plasma miR-200c-3p, miR-100-5p, and miR-1826 serve as potential diagnostic biomarkers for knee osteoarthritis: Randomized controlled trials. Medicine (Baltimore). 2019;98(51):e18110. doi:10.1097/MD.0000000000018110
69. Lambert C, Dubuc JE, Montell E, et al. Gene expression pattern of cells from inflamed and normal areas of osteoarthritis synovial membrane. Arthritis Rheumatol. 2014;66(4):960-968. doi:10.1002/art.38315
70. Lee JS, Im GI. SOX trio decrease in the articular cartilage with the advancement of osteoarthritis. Connect Tissue Res. 2011;52(6):496-502. doi:10.3109/03008207.2011.585409
71. Li C, Luo J, Xu X, et al. Single cell sequencing revealed the underlying pathogenesis of the development of osteoarthritis. Gene. 2020;757:144939. doi:10.1016/j.gene.2020.144939
72. Li F, Yao J, Hao Q, Duan Z. miRNA-103 promotes chondrocyte apoptosis by down-regulation of Sphingosine kinase-1 and ameliorates PI3K/AKT pathway in osteoarthritis. Biosci Rep. 2019;39(10):BSR20191255. doi:10.1042/BSR20191255
73. Li H, Li Z, Pi Y, et al. MicroRNA-375 exacerbates knee osteoarthritis through repressing chondrocyte autophagy by targeting ATG2B. Aging (Albany NY). 2020;12(8):7248-7261. doi:10.18632/aging.103073
74. Li HZ, Xu XH, Lin N, et al. Overexpression of miR-10a-5p facilitates the progression of osteoarthritis. Aging (Albany NY). 2020;12(7):5948-5976. doi:10.18632/aging.102989
75. Li X, Liao Z, Deng Z, Chen N, Zhao L. Combining bulk and single-cell RNA-sequencing data to reveal gene expression pattern of chondrocytes in the osteoarthritic knee. Bioengineered. 2021;12(1):997-1007. doi:10.1080/21655979.2021.1903207
76. Li YF, Li SH, Liu Y, Luo YT. Long Noncoding RNA CIR Promotes Chondrocyte Extracellular Matrix Degradation in Osteoarthritis by Acting as a Sponge For Mir-27b. Cell Physiol Biochem. 2017;43(2):602-610. doi:10.1159/000480532
77. Li Z, Yuan B, Pei Z, et al. Circ_0136474 and MMP-13 suppressed cell proliferation by competitive binding to miR-127-5p in osteoarthritis. J Cell Mol Med. 2019;23(10):6554-6564. doi:10.1111/jcmm.14400
78. Lian WS, Ko JY, Wu RW, et al. MicroRNA-128a represses chondrocyte autophagy and exacerbates knee osteoarthritis by disrupting Atg12. Cell Death Dis. 2018;9(9):919. doi:10.1038/s41419-018-0994-y
79. Liu Y, Yau MS, Yerges-Armstrong LM, et al. Genetic Determinants of Radiographic Knee Osteoarthritis in African Americans. J Rheumatol. 2017;44(11):1652-1658. doi:10.3899/jrheum.161488
80. Liu Z, Chen S, Yang Y, et al. MicroRNA 671 3p regulates the development of knee osteoarthritis by targeting TRAF3 in chondrocytes. Mol Med Rep. 2019;20(3):2843-2850. doi:10.3892/mmr.2019.10488
81. Lu M, Zhou E. Long noncoding RNA LINC00662-miR-15b-5p mediated GPR120 dysregulation contributes to osteoarthritis [published correction appears in Pathol Int. 2020 Mar;70(3):189]. Pathol Int. 2020;70(3):155-165. doi:10.1111/pin.12875
82. Lu X, Yu Y, Yin F, et al. Knockdown of PVT1 inhibits IL-1β-induced injury in chondrocytes by regulating miR-27b-3p/TRAF3 axis. Int Immunopharmacol. 2020;79:106052. doi:10.1016/j.intimp.2019.106052
83. M Dunn C, Nevitt MC, Lynch JA, Jeffries MA. A pilot study of peripheral blood DNA methylation models as predictors of knee osteoarthritis radiographic progression: data from the Osteoarthritis Initiative (OAI). Sci Rep. 2019;9(1):16880. doi:10.1038/s41598-019-53298-9
84. Ma F, Li G, Yu Y, Xu J, Wu X. MiR-33b-3p promotes chondrocyte proliferation and inhibits chondrocyte apoptosis and cartilage ECM degradation by targeting DNMT3A in osteoarthritis. Biochem Biophys Res Commun. 2019;519(2):430-437. doi:10.1016/j.bbrc.2019.09.022
85. Mao G, Hu S, Zhang Z, et al. Exosomal miR-95-5p regulates chondrogenesis and cartilage degradation via histone deacetylase 2/8. J Cell Mol Med. 2018;22(11):5354-5366. doi:10.1111/jcmm.13808
86. Miyamoto Y, Shi D, Nakajima M, et al. Common variants in DVWA on chromosome 3p24.3 are associated with susceptibility to knee osteoarthritis. Nat Genet. 2008;40(8):994-998. doi:10.1038/ng.176
87. Moazedi-Fuerst FC, Hofner M, Gruber G, et al. Epigenetic differences in human cartilage between mild and severe OA. J Orthop Res. 2014;32(12):1636-1645. doi:10.1002/jor.22722
88. Mokuda S, Nakamichi R, Matsuzaki T, et al. Wwp2 maintains cartilage homeostasis through regulation of Adamts5. Nat Commun. 2019;10(1):2429. doi:10.1038/s41467-019-10177-1
89. Nakajima M, Takahashi A, Kou I, et al. New sequence variants in HLA class II/III region associated with susceptibility to knee osteoarthritis identified by genome-wide association study. PLoS One. 2010;5(3):e9723. doi:10.1371/journal.pone.0009723
90. Nalesso G, Thorup AS, Eldridge SE, et al. Calcium calmodulin kinase II activity is required for cartilage homeostasis in osteoarthritis. Sci Rep. 2021;11(1):5682. doi:10.1038/s41598-021-82067-w
91. Nanus DE, Wijesinghe SN, Pearson MJ, et al. Regulation of the Inflammatory Synovial Fibroblast Phenotype by Metastasis-Associated Lung Adenocarcinoma Transcript 1 Long Noncoding RNA in Obese Patients With Osteoarthritis. Arthritis Rheumatol. 2020;72(4):609-619. doi:10.1002/art.41158
92. Naot D, Bentley J, Macpherson C, et al. Molecular characterisation of osteoblasts from bone obtained from people of Polynesian and European ancestry undergoing joint replacement surgery. Sci Rep. 2021;11(1):2428. doi:10.1038/s41598-021-81731-5
93. Ni Z, Kuang L, Chen H, et al. The exosome-like vesicles from osteoarthritic chondrocyte enhanced mature IL-1β production of macrophages and aggravated synovitis in osteoarthritis. Cell Death Dis. 2019;10(7):522. doi:10.1038/s41419-019-1739-2
94. Ntoumou E, Tzetis M, Braoudaki M, et al. Serum microRNA array analysis identifies miR-140-3p, miR-33b-3p and miR-671-3p as potential osteoarthritis biomarkers involved in metabolic processes. Clin Epigenetics. 2017;9:127. doi:10.1186/s13148-017-0428-1
95. Otero M, Peng H, Hachem KE, et al. ELF3 modulates type II collagen gene (COL2A1) transcription in chondrocytes by inhibiting SOX9-CBP/p300-driven histone acetyltransferase activity. Connect Tissue Res. 2017;58(1):15-26. doi:10.1080/03008207.2016.1200566
96. Papathanasiou I, Kostopoulou F, Malizos KN, Tsezou A. DNA methylation regulates sclerostin (SOST) expression in osteoarthritic chondrocytes by bone morphogenetic protein 2 (BMP-2) induced changes in Smads binding affinity to the CpG region of SOST promoter. Arthritis Res Ther. 2015;17(1):160. doi:10.1186/s13075-015-0674-6
97. Qiu WJ, Xu MZ, Zhu XD, Ji YH. MicroRNA-27a alleviates IL-1β-induced inflammatory response and articular cartilage degradation via TLR4/NF-κB signaling pathway in articular chondrocytes. Int Immunopharmacol. 2019;76:105839. doi:10.1016/j.intimp.2019.105839
98. Ramos YF, Bos SD, Lakenberg N, et al. Genes expressed in blood link osteoarthritis with apoptotic pathways. Ann Rheum Dis. 2014;73(10):1844-1853. doi:10.1136/annrheumdis-2013-203405
99. Ren T, Wei P, Song Q, Ye Z, Wang Y, Huang L. MiR-140-3p Ameliorates the Progression of Osteoarthritis via Targeting CXCR4. Biol Pharm Bull. 2020;43(5):810-816. doi:10.1248/bpb.b19-00959
100. Rerknimitr P, Kantikosum K, Chottawornsak N, et al. Chronic occupational exposure to lead leads to significant mucocutaneous changes in lead factory workers. J Eur Acad Dermatol Venereol. 2019;33(10):1993-2000. doi:10.1111/jdv.15678
101. Reynard LN, Bui C, Syddall CM, Loughlin J. CpG methylation regulates allelic expression of GDF5 by modulating binding of SP1 and SP3 repressor proteins to the osteoarthritis susceptibility SNP rs143383. Hum Genet. 2014;133(8):1059-1073. doi:10.1007/s00439-014-1447-z
102. Rice SJ, Aubourg G, Sorial AK, et al. Identification of a novel, methylation-dependent, RUNX2 regulatory region associated with osteoarthritis risk. Hum Mol Genet. 2018;27(19):3464-3474. doi:10.1093/hmg/ddy257
103. Rice SJ, Tselepi M, Sorial AK, et al. Prioritization of PLEC and GRINA as Osteoarthritis Risk Genes Through the Identification and Characterization of Novel Methylation Quantitative Trait Loci. Arthritis Rheumatol. 2019;71(8):1285-1296. doi:10.1002/art.40849
104. Rousseau JC, Millet M, Croset M, Sornay-Rendu E, Borel O, Chapurlat R. Association of circulating microRNAs with prevalent and incident knee osteoarthritis in women: the OFELY study. Arthritis Res Ther. 2020;22(1):2. doi:10.1186/s13075-019-2086-5
105. Rushton MD, Reynard LN, Barter MJ, et al. Characterization of the cartilage DNA methylome in knee and hip osteoarthritis. Arthritis Rheumatol. 2014;66(9):2450-2460. doi:10.1002/art.38713
106. Rushton MD, Reynard LN, Young DA, et al. Methylation quantitative trait locus analysis of osteoarthritis links epigenetics with genetic risk. Hum Mol Genet. 2015;24(25):7432-7444. doi:10.1093/hmg/ddv433
107. Rushton MD, Young DA, Loughlin J, Reynard LN. Differential DNA methylation and expression of inflammatory and zinc transporter genes defines subgroups of osteoarthritic hip patients. Ann Rheum Dis. 2015;74(9):1778-1782. doi:10.1136/annrheumdis-2014-206752
108. Scott JL, Gabrielides C, Davidson RK, et al. Superoxide dismutase downregulation in osteoarthritis progression and end-stage disease [published correction appears in Ann Rheum Dis. 2011 Feb;70(2):397. Talyor, Robert W [corrected to Taylor, Robert W]]. Ann Rheum Dis. 2010;69(8):1502-1510. doi:10.1136/ard.2009.119966
109. Seidl CI, Murphy CL. Dual and Opposing Regulation of MMP1 and MMP13 by Both Arms of miR-675 in Human Articular Chondrocytes. Cell Physiol Biochem. 2019;53(1):172-185. doi:10.33594/000000128
110. Shen P, Yang Y, Liu G, et al. CircCDK14 protects against Osteoarthritis by sponging miR-125a-5p and promoting the expression of Smad2. Theranostics. 2020;10(20):9113-9131. doi:10.7150/thno.45993
111. Shi J, Guo K, Su S, Li J, Li C. miR 486 5p is upregulated in osteoarthritis and inhibits chondrocyte proliferation and migration by suppressing SMAD2. Mol Med Rep. 2018;18(1):502-508. doi:10.3892/mmr.2018.8931
112. Shu Y, Long J, Guo W, Ye W. MicroRNA 195 5p inhibitor prevents the development of osteoarthritis by targeting REGγ. Mol Med Rep. 2019;19(6):4561-4568. doi:10.3892/mmr.2019.10124
113. Skrzypa M, Szala D, Gablo N, et al. miRNA-146a-5p is upregulated in serum and cartilage samples of patients with osteoarthritis. Polish Journal of Surgery. 2019;91: 1-5. doi: 10.5604/01.3001.0013.0135
114. Song X, Zhu M, Sun Y, Liu B, Yan Z, Yin Y. MiR-204 enhances the progression of osteoarthritis by suppressing the production of IL-1β. Pharmazie. 2017;72(10):587-592. doi:10.1691/ph.2017.7588
115. Sorial AK, Hofer IMJ, Tselepi M, et al. Multi-tissue epigenetic analysis of the osteoarthritis susceptibility locus mapping to the plectin gene PLEC. Osteoarthritis Cartilage. 2020;28(11):1448-1458. doi:10.1016/j.joca.2020.06.001
116. Steinberg J, Ritchie GRS, Roumeliotis TI, et al. Integrative epigenomics, transcriptomics and proteomics of patient chondrocytes reveal genes and pathways involved in osteoarthritis. Sci Rep. 2017;7(1):8935. doi:10.1038/s41598-017-09335-6
117. Styrkarsdottir U, Helgason H, Sigurdsson A, et al. Whole-genome sequencing identifies rare genotypes in COMP and CHADL associated with high risk of hip osteoarthritis [published correction appears in Nat Genet. 2017 Jul 27;49(8):1286]. Nat Genet. 2017;49(5):801-805. doi:10.1038/ng.3816
118. Styrkarsdottir U, Lund SH, Thorleifsson G, et al. Meta-analysis of Icelandic and UK data sets identifies missense variants in SMO, IL11, COL11A1 and 13 more new loci associated with osteoarthritis. Nat Genet. 2018;50(12):1681-1687. doi:10.1038/s41588-018-0247-0
119. Styrkarsdottir U, Thorleifsson G, Helgadottir HT, et al. Severe osteoarthritis of the hand associates with common variants within the ALDH1A2 gene and with rare variants at 1p31. Nat Genet. 2014;46(5):498-502. doi:10.1038/ng.2957
120. Sun H, Wen X, Li H, et al. Single-cell RNA-seq analysis identifies meniscus progenitors and reveals the progression of meniscus degeneration. Ann Rheum Dis. 2020;79(3):408-417. doi:10.1136/annrheumdis-2019-215926
121. Sun Y, Mauerhan DR, Honeycutt PR, et al. Analysis of meniscal degeneration and meniscal gene expression. BMC Musculoskelet Disord. 2010;11:19. doi:10.1186/1471-2474-11-19
122. Tachmazidou I, Hatzikotoulas K, Southam L, et al. Identification of new therapeutic targets for osteoarthritis through genome-wide analyses of UK Biobank data. Nat Genet. 2019;51(2):230-236. doi:10.1038/s41588-018-0327-1
123. Takahashi A, de Andrés MC, Hashimoto K, et al. DNA methylation of the RUNX2 P1 promoter mediates MMP13 transcription in chondrocytes. Sci Rep. 2017;7(1):7771. doi:10.1038/s41598-017-08418-8
124. Takahashi A, de Andrés MC, Hashimoto K, Itoi E, Oreffo RO. Epigenetic regulation of interleukin-8, an inflammatory chemokine, in osteoarthritis. Osteoarthritis Cartilage. 2015;23(11):1946-1954. doi:10.1016/j.joca.2015.02.168
125. Tian F, Wang J, Zhang Z, Yang J. LncRNA SNHG7/miR-34a-5p/SYVN1 axis plays a vital role in proliferation, apoptosis and autophagy in osteoarthritis. Biol Res. 2020;53(1):9. doi:10.1186/s40659-020-00275-6
126. Tu Y, Ma T, Wen T, et al. MicroRNA-377-3p alleviates IL-1β-caused chondrocyte apoptosis and cartilage degradation in osteoarthritis in part by downregulating ITGA6. Biochem Biophys Res Commun. 2020;523(1):46-53. doi:10.1016/j.bbrc.2019.11.186
127. Valdes AM, Evangelou E, Kerkhof HJ, et al. The GDF5 rs143383 polymorphism is associated with osteoarthritis of the knee with genome-wide statistical significance. Ann Rheum Dis. 2011;70(5):873-875. doi:10.1136/ard.2010.134155
128. Wang H, Zhang H, Sun Q, et al. Intra-articular Delivery of Antago-miR-483-5p Inhibits Osteoarthritis by Modulating Matrilin 3 and Tissue Inhibitor of Metalloproteinase 2. Mol Ther. 2017;25(3):715-727. doi:10.1016/j.ymthe.2016.12.020
129. Wang J, Fang L, Ye L, et al. miR-137 targets the inhibition of TCF4 to reverse the progression of osteoarthritis through the AMPK/NF-κB signaling pathway. Biosci Rep. 2020;40(6):BSR20200466. doi:10.1042/BSR20200466
130. Wang J, Wang Y, Zhang H, et al. Forkhead box C1 promotes the pathology of osteoarthritis by upregulating β-catenin in synovial fibroblasts. FEBS J. 2020;287(14):3065-3087. doi:10.1111/febs.15178
131. Wang P, Dong R, Wang B, et al. Genome-wide microRNA screening reveals miR-582-5p as a mesenchymal stem cell-specific microRNA in subchondral bone of the human knee joint. J Cell Physiol. 2019;234(12):21877-21888. doi:10.1002/jcp.28751
132. Wang S, Zheng Y, Hu Z, Wang Z, Zhang Y, Wei L. Downregulated miR 302d 3p promotes chondrocyte proliferation and migration by regulation of Unc 51 like kinase 1. Int J Mol Med. 2019;44(3):1039-1047. doi:10.3892/ijmm.2019.4267
133. Wang T, Hao Z, Liu C, et al. LEF1 mediates osteoarthritis progression through circRNF121/miR-665/MYD88 axis via NF-кB signaling pathway [published correction appears in Cell Death Dis. 2020 Aug 11;11(8):689]. Cell Death Dis. 2020;11(7):598. doi:10.1038/s41419-020-02769-3
134. Wang X, Ning Y, Zhang P, et al. Comparison of the major cell populations among osteoarthritis, Kashin-Beck disease and healthy chondrocytes by single-cell RNA-seq analysis. Cell Death Dis. 2021;12(6):551. doi:10.1038/s41419-021-03832-3
135. Wang Y, Shen S, Li Z, Li W, Weng X. MIR-140-5p affects chondrocyte proliferation, apoptosis, and inflammation by targeting HMGB1 in osteoarthritis. Inflamm Res. 2020;69(1):63-73. doi:10.1007/s00011-019-01294-0
136. Wang Z, Hu J, Pan Y, et al. miR-140-5p/miR-149 Affects Chondrocyte Proliferation, Apoptosis, and Autophagy by Targeting FUT1 in Osteoarthritis [published correction appears in Inflammation. 2019 Aug;42(4):1515-1516]. Inflammation. 2018;41(3):959-971. doi:10.1007/s10753-018-0750-6
137. Wang Z, Li X, Yang J, et al. Single-cell RNA sequencing deconvolutes the in vivo heterogeneity of human bone marrow-derived mesenchymal stem cells. bioRxiv, 2020. doi:10.1101/2020.04.06.027904
138. Wu J, Zou M, Ping A, Deng Z, Cai L. MicroRNA-449a upregulation promotes chondrocyte extracellular matrix degradation in osteoarthritis. Biomed Pharmacother. 2018;105:940-946. doi:10.1016/j.biopha.2018.06.074
139. Wu Z, Shou L, Wang J, Xu X. Identification of the key gene and pathways associated with osteoarthritis via single-cell RNA sequencing on synovial fibroblasts. Medicine (Baltimore). 2020;99(33):e21707. doi:10.1097/MD.0000000000021707
140. Xu K, Meng Z, Xian XM, et al. LncRNA PVT1 induces chondrocyte apoptosis through upregulation of TNF-α in synoviocytes by sponging miR-211-3p. Mol Cell Probes. 2020;52:101560. doi:10.1016/j.mcp.2020.101560
141. Xue H, Tu Y, Ma T, et al. miR-93-5p attenuates IL-1β-induced chondrocyte apoptosis and cartilage degradation in osteoarthritis partially by targeting TCF4. Bone. 2019;123:129-136. doi:10.1016/j.bone.2019.03.035
142. Xue J, Min Z, Xia Z, et al. The hsa-miR-181a-5p reduces oxidation resistance by controlling SECISBP2 in osteoarthritis. BMC Musculoskelet Disord. 2018;19(1):355. doi:10.1186/s12891-018-2273-6
143. Yang F, Zhou S, Wang C, et al. Epigenetic modifications of interleukin-6 in synovial fibroblasts from osteoarthritis patients. Sci Rep. 2017;7:43592. doi:10.1038/srep43592
144. Yang J, Wang N. Genome-wide expression and methylation profiles reveal candidate genes and biological processes underlying synovial inflammatory tissue of patients with osteoarthritis. Int J Rheum Dis. 2015;18(7):783-790. doi:10.1111/1756-185X.12643
145. Yang Q, Zhou Y, Cai P, et al. Downregulation of microRNA-23b-3p alleviates IL-1β-induced injury in chondrogenic CHON-001 cells. Drug Des Devel Ther. 2019;13:2503-2512. doi:10.2147/DDDT.S211051
146. Yang Y, Xing D, Wang Y, Jia H, Li B, Li JJ. A long non-coding RNA, HOTAIR, promotes cartilage degradation in osteoarthritis by inhibiting WIF-1 expression and activating Wnt pathway. BMC Mol Cell Biol. 2020;21(1):53. doi:10.1186/s12860-020-00299-6
147. Yi P, Xu X, Yao J, Qiu B. Analysis of mRNA Expression and DNA Methylation Datasets According to the Genomic Distribution of CpG Sites in Osteoarthritis. Front Genet. 2021;12:618803. doi:10.3389/fgene.2021.618803
148. Zengini E, Hatzikotoulas K, Tachmazidou I, et al. Genome-wide analyses using UK Biobank data provide insights into the genetic architecture of osteoarthritis. Nat Genet. 2018;50(4):549-558. doi:10.1038/s41588-018-0079-y
149. Zerrouk N, Miagoux Q, Dispot A, Elati M, Niarakis A. Identification of putative master regulators in rheumatoid arthritis synovial fibroblasts using gene expression data and network inference. Sci Rep. 2020;10(1):16236. doi:10.1038/s41598-020-73147-4
150. Zhang C, Zhang Z, Chang Z, et al. miR-193b-5p regulates chondrocytes metabolism by directly targeting histone deacetylase 7 in interleukin-1β-induced osteoarthritis. J Cell Biochem. 2019;120(8):12775-12784. doi:10.1002/jcb.28545
151. Zhang L, Yang M, Marks P, et al. Serum non-coding RNAs as biomarkers for osteoarthritis progression after ACL injury. Osteoarthritis Cartilage. 2012;20(12):1631-1637. doi:10.1016/j.joca.2012.08.016
152. Zhang X, Huang CR, Pan S, et al. Long non-coding RNA SNHG15 is a competing endogenous RNA of miR-141-3p that prevents osteoarthritis progression by upregulating BCL2L13 expression. Int Immunopharmacol. 2020;83:106425. doi:10.1016/j.intimp.2020.106425
153. Zhang X, Huang N, Huang R, et al. Single-cell rna-seq analysis identifies the biomarkers and differentiation of chondrocyte in human osteoarthritis. Am J Transl Res. 2020;12(11):7326-7339.
154. Zhang Y, Fukui N, Yahata M, et al. Genome-wide DNA methylation profile implicates potential cartilage regeneration at the late stage of knee osteoarthritis. Osteoarthritis Cartilage. 2016;24(5):835-843. doi:10.1016/j.joca.2015.12.013
155. Zhao L, Wang Q, Zhang C, Huang C. Genome-wide DNA methylation analysis of articular chondrocytes identifies TRAF1, CTGF, and CX3CL1 genes as hypomethylated in osteoarthritis. Clin Rheumatol. 2017;36(10):2335-2342. doi:10.1007/s10067-017-3667-9
156. Zhao X, Li H, Wang L. MicroRNA-107 regulates autophagy and apoptosis of osteoarthritis chondrocytes by targeting TRAF3. Int Immunopharmacol. 2019;71:181-187. doi:10.1016/j.intimp.2019.03.005
157. Zheng W D, Zhou F L, Lin N, et al. Investigation for the role of CTX-III and microRNA-98 in diagnosis and treatment of osteoarthritis. Eur Rev Med Pharmacol Sci. 2018;22(17): 5424-5428. doi:10.26355/eurrev_201809_15801
158. Zhou J, Zhao Z, He C, et al. Single-Cell Transcriptome Analysis Profile of Meniscal Tissue Macrophages in Human Osteoarthritis. J Immunol Res. 2020;2020:8127281. doi:10.1155/2020/8127281
159. Zhu H, Hu Y, Wang C, Zhang X, He D. CircGCN1L1 promotes synoviocyte proliferation and chondrocyte apoptosis by targeting miR-330-3p and TNF-α in TMJ osteoarthritis. Cell Death Dis. 2020;11(4):284. doi:10.1038/s41419-020-2447-7